Main Article Content
Abstract
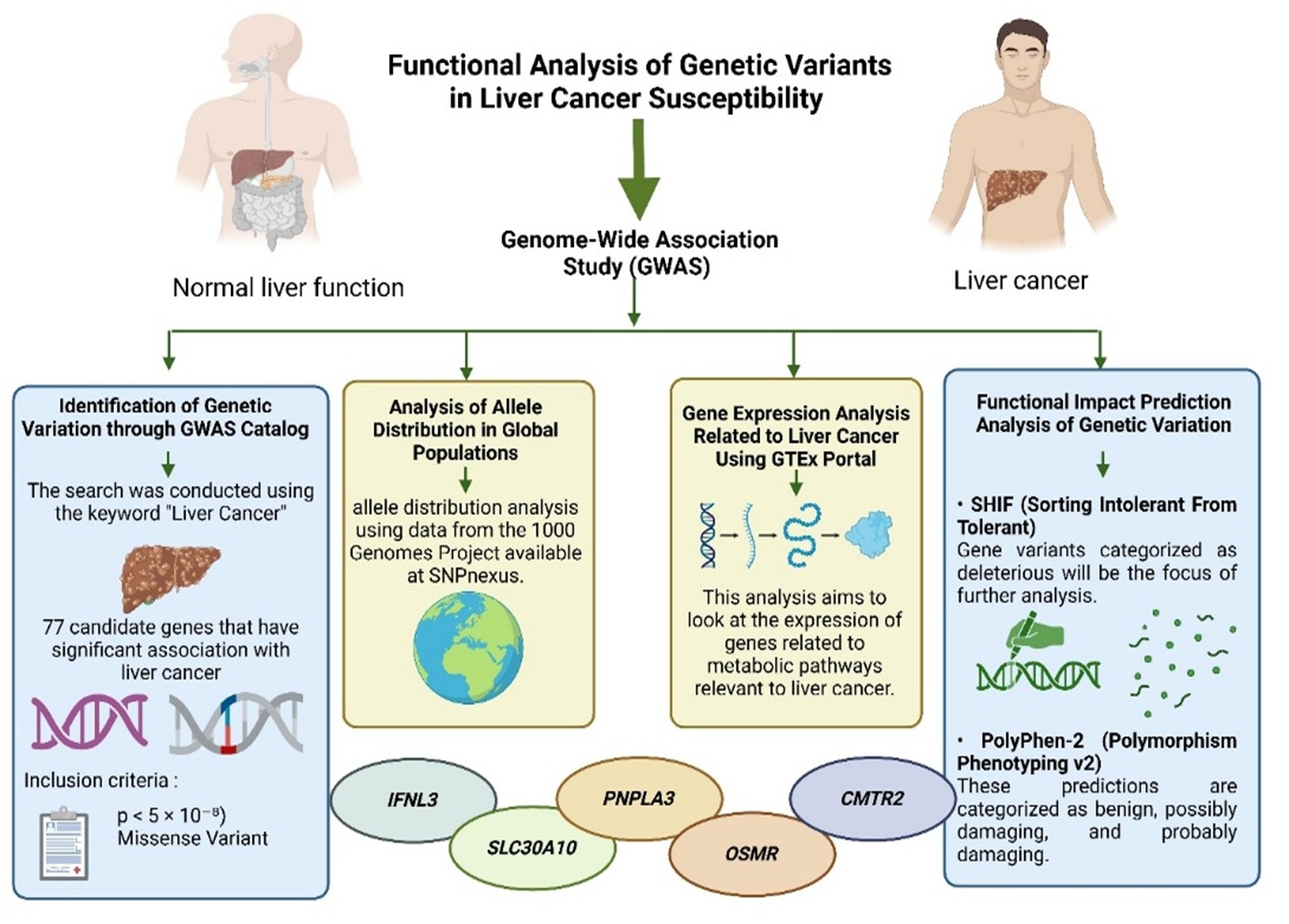
Liver cancer is one of the leading causes of cancer death worldwide. Genetic factors play a role in determining a person's susceptibility to this disease. Genome-Wide Association Studies (GWAS) have identified several genetic variants associated with liver cancer, but their functional mechanisms still need to be further explored. Therefore, this study aims to identify genetic variants that contribute to liver cancer, evaluate their functional effects on proteins, analyze allele frequencies across global populations, and examine gene expression in various human tissues. This study used a bioinformatics approach to identify genetic variations associated with liver cancer from the GWAS Catalog. Five selected missense variants were analyzed using SIFT and PolyPhen-2 to assess their functional impact. Allele distributions in the global population were analyzed using 1000 Genomes Project data, and gene expression was analyzed using the GTEx Portal. The analysis identified 77 candidate genes with significant associations with liver cancer, based on p-values meeting the threshold (p < 5 × 10⁻⁸). Five genes in the Missense Variant category showed a strong association with liver cancer: IFNL3, SLC30A10, PNPLA3, OSMR, and CMTR2. In the analysis using SIFT and PolyPhen-2, the rs3096380 variant (CMTR2) was deleterious, and rs738409 (PNPLA3) and rs188273166 (SLC30A10) were deleterious and probably damaging, with the potential to disrupt protein function and contribute to the pathogenesis of liver cancer. Conclusion: Genetic variations rs738409 (PNPLA3) and rs188273166 (SLC30A10) are deleterious and probably damaging, potentially disrupting protein function and contributing to liver cancer pathogenesis.
Article Details

This work is licensed under a Creative Commons Attribution 4.0 International License.
References
- Amanzada, A., Reinhardt, L., Fey, D., Zeisberg, E. M., & Mihm, S. (2015). Hepatic Interferon-λ3 (IFNL3) Gene Expression Reveals Not to Be Attenuated in Non-Favorable IFNL3 rs4803217 or IFNL4 rs368234815 Minor Allele Carriers in Chronic Hepatitis C. PLoS ONE, 10(11). https://doi.org/10.1371/journal.pone.0143783
- Amukti, D. P., Irham, L. M., Surono, S., Adikusuma, W., Chong, R., El Khair, R., Purwanto, B. D., Satria, R. D., Ates, I., Khairi, S., & Rungprai, D. (2024). Genomic Variants And Epidemiology Of Atherosclerosis – Worldwide Correlation Analysis And Utilization Of Atherosclerosis Gene Variants For Identification Of Drug Target Candidates With Bioinformatics Approach. Bulgarian Cardiology, 30(4), 108–120. https://doi.org/10.3897/bgcardio.30.e145391
- Ashtari, S., Pourhoseingholi, M. A., Sharifian, A., & Zali, M. R. (2015). Hepatocellular carcinoma in Asia: Prevention strategy and planning. In World Journal of Hepatology (Vol. 7, Issue 12, pp. 1708–1717). Baishideng Publishing Group Co. https://doi.org/10.4254/wjh.v7.i12.1708
- Bharti, N., Banerjee, R., Achalare, A., Kasibhatla, S. M., & Joshi, R. (2024). Estimation of genetic variation in vitiligo associated genes: Population genomics perspective. BMC Genomic Data, 25(1). https://doi.org/10.1186/s12863-024-01254-6
- Byrne, K. T., Vonderheide, R. H., Jaffee, E. M., & Armstrong, T. D. (2015). Special conference on tumor immunology and immunotherapy: A new chapter. Cancer Immunology Research, 3(6), 590–597. https://doi.org/10.1158/2326-6066.CIR-15-0106
- Cao, W., Chen, H. Da, Yu, Y. W., Li, N., & Chen, W. Q. (2021). Changing profiles of cancer burden worldwide and in China: A secondary analysis of the global cancer statistics 2020. Chinese Medical Journal, 134(7), 783–791. https://doi.org/10.1097/CM9.0000000000001474
- Cao, W., Qin, K., Li, F., & Chen, W. (2024). Comparative study of cancer profiles between 2020 and 2022 using global cancer statistics (GLOBOCAN). Journal of the National Cancer Center, 4(2), 128–134. https://doi.org/10.1016/j.jncc.2024.05.001
- Chatziparasidou, A., Kyrgiafini, M. A., Sarafidou, T., Moutou, K. A., & Mamuris, Z. (2024). Genetic Insights into Azoospermia and Severe Oligozoospermia: Discovering Seven SNPs through GWAS and In Silico Analysis. Current Issues in Molecular Biology, 46(7), 6522–6532. https://doi.org/10.3390/cimb46070389
- Fan, S., Zhao, T., & Sun, L. (2023). The global prevalence and ethnic heterogeneity of iron-refractory iron deficiency anaemia. Orphanet Journal of Rare Diseases, 18(1). https://doi.org/10.1186/s13023-022-02612-2
- Gumelar, G., Ulfa, M. M., Amukti, D. P., Irham, L. M., Yuliani, S., Adikusuma, W., Khairi, S., Darmawi, D., Chong, R., Ates, I., Singh, D., & Chavan, A. A. (2024). Harnessing Genomic and Bioinformatic Data to Broaden Understanding of Leukaemia Across Continents. Scripta Medica (Banja Luka), 55(6), 717–725. https://doi.org/10.5937/scriptamed55-51720
- Guo, Q., Zhu, X., Beeraka, N. M., Zhao, R., Li, S., Li, F., Mahesh, P. A., Nikolenko, V. N., Fan, R., & Liu, J. (2024). Projected epidemiological trends and burden of liver cancer by 2040 based on GBD, CI5plus, and WHO data. Scientific Reports, 14(1). https://doi.org/10.1038/s41598-024-77658-2
- Hewitt, D. B., Brown, Z. J., & Pawlik, T. M. (2022). Current Perspectives on the Surgical Management of Perihilar Cholangiocarcinoma. In Cancers (Vol. 14, Issue 9). MDPI. https://doi.org/10.3390/cancers14092208
- Humolungo, D. T. W. S., Anjani, R., Irham, L. M., Sulistyani, N., Ma’ruf, M., Amukti, D. P., Adikusuma, W., Sarasmita, M. A., Khairi, S., Purwanto, B. D., Suyatmi, S., Siswanto, L. M. H., Satria, R. D., Pranata, S., & Chong, R. (2024). Identification of pathogenic gene variants in carpal tunnel syndrome using bioinformatics approaches. E3S Web of Conferences, 501. https://doi.org/10.1051/e3sconf/202450101022
- Ignatochkina, A. V., Iguchi, J. A., Kore, A. R., & Ho, C. K. (2024). Trypanosome mRNA recapping is triggered by hypermethylation originating from cap 4. Nucleic Acids Research. https://doi.org/10.1093/nar/gkae614
- Inesta-Vaquera, F., & Cowling, V. H. (2017). Regulation and function of CMTR1-dependent mRNA cap methylation. In Wiley Interdisciplinary Reviews: RNA (Vol. 8, Issue 6). Blackwell Publishing Ltd. https://doi.org/10.1002/wrna.1450
- Linskey, D. W., Linskey, D. C., McLeod, H. L., & Luzum, J. A. (2021). The need to shift pharmacogenetic research from candidate gene to genome-wide association studies. Pharmacogenomics, 22(17), 1143–1150. https://doi.org/10.2217/pgs-2021-0108
- Liu, M., & Park, S. (2024). The Role of PNPLA3_rs738409 Gene Variant, Lifestyle Factors, and Bioactive Compounds in Nonalcoholic Fatty Liver Disease: A Population-Based and Molecular Approach towards Healthy Nutrition. Nutrients, 16(8). https://doi.org/10.3390/nu16081239
- Manduchi, E., Orzechowski, P. R., Ritchie, M. D., & Moore, J. H. (2019). Exploration of a diversity of computational and statistical measures of association for genome-wide genetic studies. BioData Mining, 12(1). https://doi.org/10.1186/s13040-019-0201-4
- McGlynn, K. A., Petrick, J. L., & El-Serag, H. B. (2021). Epidemiology of Hepatocellular Carcinoma. In Hepatology (Vol. 73, Issue S1, pp. 4–13). John Wiley and Sons Inc. https://doi.org/10.1002/hep.31288
- Mercadante, C. J., Prajapati, M., Conboy, H. L., Dash, M. E., Herrera, C., Pettiglio, M. A., Cintron-Rivera, L., Salesky, M. A., Rao, D. B., & Bartnikas, T. B. (2019). Manganese transporter Slc30a10 controls physiological manganese excretion and toxicity. Journal of Clinical Investigation, 129(12), 5442–5461. https://doi.org/10.1172/JCI129710
- Miao, Y., Chen, Y., & Mi, D. (2022). Role of gasdermin family proteins in the occurrence and progression of hepatocellular carcinoma. In Heliyon (Vol. 8, Issue 10). Elsevier Ltd. https://doi.org/10.1016/j.heliyon.2022.e11035
- Ohi, K., Shimada, T., Yasuyama, T., Uehara, T., & Kawasaki, Y. (2017). Variability of 128 schizophrenia-associated gene variants across distinct ethnic populations. Translational Psychiatry, 7(1). https://doi.org/10.1038/tp.2016.260
- Pratiwi, N., Ulfah, A. J., Rachmadina, R., Irham, L. M., Afief, A. R., Adikusuma, W., Darmawi, D., Kemal, R. A., Rangkuti, I. F., & Savira, M. (2024). Promising candidate drug target genes for repurposing in cervical cancer: A bioinformatics-based approach. Narra J, 4(3). https://doi.org/10.52225/narra.v4i3.938
- Rady, B., Nishio, T., Dhar, D., Liu, X., Erion, M., Kisseleva, T., Brenner, D. A., & Pocai, A. (2021). PNPLA3 downregulation exacerbates the fibrotic response in human hepatic stellate cells. PLoS ONE, 16(12 December). https://doi.org/10.1371/journal.pone.0260721
- Togninalli, M., Seren, Ü., Meng, D., Fitz, J., Nordborg, M., Weigel, D., Borgwardt, K., Korte, A., & Grimm, D. G. (2018). The AraGWAS Catalog: A curated and standardized Arabidopsis thaliana GWAS catalog. Nucleic Acids Research, 46(D1), D1150–D1156. https://doi.org/10.1093/nar/gkx954
- Yang, Z., Zhang, T., Han, S., Kusumanchi, P., Huda, N., Jiang, Y., & Liangpunsakul, S. (2021). Long noncoding RNA H19 – a new player in the pathogenesis of liver diseases. In Translational Research (Vol. 230, pp. 139–150). Mosby Inc. https://doi.org/10.1016/j.trsl.2020.11.010
- Zhang, T., Hu, Y., Wu, X., Ma, R., Jiang, Q., & Wang, Y. (2016). Identifying Liver Cancer-Related Enhancer SNPs by Integrating GWAS and Histone Modification ChIP-seq Data. BioMed Research International, 2016. https://doi.org/10.1155/2016/2395341
- Zhu, Y. X., Li, C. H., Li, G., Feng, H., Xia, T., Wong, C. H., Fung, F. K. C., Tong, J. H. M., To, K. F., Chen, R., & Chen, Y. (2020). LLGL1 Regulates Gemcitabine Resistance by Modulating the ERK-SP1-OSMR Pathway in Pancreatic Ductal Adenocarcinoma. CMGH, 10(4), 811–828. https://doi.org/10.1016/j.jcmgh.2020.06.009
References
Amanzada, A., Reinhardt, L., Fey, D., Zeisberg, E. M., & Mihm, S. (2015). Hepatic Interferon-λ3 (IFNL3) Gene Expression Reveals Not to Be Attenuated in Non-Favorable IFNL3 rs4803217 or IFNL4 rs368234815 Minor Allele Carriers in Chronic Hepatitis C. PLoS ONE, 10(11). https://doi.org/10.1371/journal.pone.0143783
Amukti, D. P., Irham, L. M., Surono, S., Adikusuma, W., Chong, R., El Khair, R., Purwanto, B. D., Satria, R. D., Ates, I., Khairi, S., & Rungprai, D. (2024). Genomic Variants And Epidemiology Of Atherosclerosis – Worldwide Correlation Analysis And Utilization Of Atherosclerosis Gene Variants For Identification Of Drug Target Candidates With Bioinformatics Approach. Bulgarian Cardiology, 30(4), 108–120. https://doi.org/10.3897/bgcardio.30.e145391
Ashtari, S., Pourhoseingholi, M. A., Sharifian, A., & Zali, M. R. (2015). Hepatocellular carcinoma in Asia: Prevention strategy and planning. In World Journal of Hepatology (Vol. 7, Issue 12, pp. 1708–1717). Baishideng Publishing Group Co. https://doi.org/10.4254/wjh.v7.i12.1708
Bharti, N., Banerjee, R., Achalare, A., Kasibhatla, S. M., & Joshi, R. (2024). Estimation of genetic variation in vitiligo associated genes: Population genomics perspective. BMC Genomic Data, 25(1). https://doi.org/10.1186/s12863-024-01254-6
Byrne, K. T., Vonderheide, R. H., Jaffee, E. M., & Armstrong, T. D. (2015). Special conference on tumor immunology and immunotherapy: A new chapter. Cancer Immunology Research, 3(6), 590–597. https://doi.org/10.1158/2326-6066.CIR-15-0106
Cao, W., Chen, H. Da, Yu, Y. W., Li, N., & Chen, W. Q. (2021). Changing profiles of cancer burden worldwide and in China: A secondary analysis of the global cancer statistics 2020. Chinese Medical Journal, 134(7), 783–791. https://doi.org/10.1097/CM9.0000000000001474
Cao, W., Qin, K., Li, F., & Chen, W. (2024). Comparative study of cancer profiles between 2020 and 2022 using global cancer statistics (GLOBOCAN). Journal of the National Cancer Center, 4(2), 128–134. https://doi.org/10.1016/j.jncc.2024.05.001
Chatziparasidou, A., Kyrgiafini, M. A., Sarafidou, T., Moutou, K. A., & Mamuris, Z. (2024). Genetic Insights into Azoospermia and Severe Oligozoospermia: Discovering Seven SNPs through GWAS and In Silico Analysis. Current Issues in Molecular Biology, 46(7), 6522–6532. https://doi.org/10.3390/cimb46070389
Fan, S., Zhao, T., & Sun, L. (2023). The global prevalence and ethnic heterogeneity of iron-refractory iron deficiency anaemia. Orphanet Journal of Rare Diseases, 18(1). https://doi.org/10.1186/s13023-022-02612-2
Gumelar, G., Ulfa, M. M., Amukti, D. P., Irham, L. M., Yuliani, S., Adikusuma, W., Khairi, S., Darmawi, D., Chong, R., Ates, I., Singh, D., & Chavan, A. A. (2024). Harnessing Genomic and Bioinformatic Data to Broaden Understanding of Leukaemia Across Continents. Scripta Medica (Banja Luka), 55(6), 717–725. https://doi.org/10.5937/scriptamed55-51720
Guo, Q., Zhu, X., Beeraka, N. M., Zhao, R., Li, S., Li, F., Mahesh, P. A., Nikolenko, V. N., Fan, R., & Liu, J. (2024). Projected epidemiological trends and burden of liver cancer by 2040 based on GBD, CI5plus, and WHO data. Scientific Reports, 14(1). https://doi.org/10.1038/s41598-024-77658-2
Hewitt, D. B., Brown, Z. J., & Pawlik, T. M. (2022). Current Perspectives on the Surgical Management of Perihilar Cholangiocarcinoma. In Cancers (Vol. 14, Issue 9). MDPI. https://doi.org/10.3390/cancers14092208
Humolungo, D. T. W. S., Anjani, R., Irham, L. M., Sulistyani, N., Ma’ruf, M., Amukti, D. P., Adikusuma, W., Sarasmita, M. A., Khairi, S., Purwanto, B. D., Suyatmi, S., Siswanto, L. M. H., Satria, R. D., Pranata, S., & Chong, R. (2024). Identification of pathogenic gene variants in carpal tunnel syndrome using bioinformatics approaches. E3S Web of Conferences, 501. https://doi.org/10.1051/e3sconf/202450101022
Ignatochkina, A. V., Iguchi, J. A., Kore, A. R., & Ho, C. K. (2024). Trypanosome mRNA recapping is triggered by hypermethylation originating from cap 4. Nucleic Acids Research. https://doi.org/10.1093/nar/gkae614
Inesta-Vaquera, F., & Cowling, V. H. (2017). Regulation and function of CMTR1-dependent mRNA cap methylation. In Wiley Interdisciplinary Reviews: RNA (Vol. 8, Issue 6). Blackwell Publishing Ltd. https://doi.org/10.1002/wrna.1450
Linskey, D. W., Linskey, D. C., McLeod, H. L., & Luzum, J. A. (2021). The need to shift pharmacogenetic research from candidate gene to genome-wide association studies. Pharmacogenomics, 22(17), 1143–1150. https://doi.org/10.2217/pgs-2021-0108
Liu, M., & Park, S. (2024). The Role of PNPLA3_rs738409 Gene Variant, Lifestyle Factors, and Bioactive Compounds in Nonalcoholic Fatty Liver Disease: A Population-Based and Molecular Approach towards Healthy Nutrition. Nutrients, 16(8). https://doi.org/10.3390/nu16081239
Manduchi, E., Orzechowski, P. R., Ritchie, M. D., & Moore, J. H. (2019). Exploration of a diversity of computational and statistical measures of association for genome-wide genetic studies. BioData Mining, 12(1). https://doi.org/10.1186/s13040-019-0201-4
McGlynn, K. A., Petrick, J. L., & El-Serag, H. B. (2021). Epidemiology of Hepatocellular Carcinoma. In Hepatology (Vol. 73, Issue S1, pp. 4–13). John Wiley and Sons Inc. https://doi.org/10.1002/hep.31288
Mercadante, C. J., Prajapati, M., Conboy, H. L., Dash, M. E., Herrera, C., Pettiglio, M. A., Cintron-Rivera, L., Salesky, M. A., Rao, D. B., & Bartnikas, T. B. (2019). Manganese transporter Slc30a10 controls physiological manganese excretion and toxicity. Journal of Clinical Investigation, 129(12), 5442–5461. https://doi.org/10.1172/JCI129710
Miao, Y., Chen, Y., & Mi, D. (2022). Role of gasdermin family proteins in the occurrence and progression of hepatocellular carcinoma. In Heliyon (Vol. 8, Issue 10). Elsevier Ltd. https://doi.org/10.1016/j.heliyon.2022.e11035
Ohi, K., Shimada, T., Yasuyama, T., Uehara, T., & Kawasaki, Y. (2017). Variability of 128 schizophrenia-associated gene variants across distinct ethnic populations. Translational Psychiatry, 7(1). https://doi.org/10.1038/tp.2016.260
Pratiwi, N., Ulfah, A. J., Rachmadina, R., Irham, L. M., Afief, A. R., Adikusuma, W., Darmawi, D., Kemal, R. A., Rangkuti, I. F., & Savira, M. (2024). Promising candidate drug target genes for repurposing in cervical cancer: A bioinformatics-based approach. Narra J, 4(3). https://doi.org/10.52225/narra.v4i3.938
Rady, B., Nishio, T., Dhar, D., Liu, X., Erion, M., Kisseleva, T., Brenner, D. A., & Pocai, A. (2021). PNPLA3 downregulation exacerbates the fibrotic response in human hepatic stellate cells. PLoS ONE, 16(12 December). https://doi.org/10.1371/journal.pone.0260721
Togninalli, M., Seren, Ü., Meng, D., Fitz, J., Nordborg, M., Weigel, D., Borgwardt, K., Korte, A., & Grimm, D. G. (2018). The AraGWAS Catalog: A curated and standardized Arabidopsis thaliana GWAS catalog. Nucleic Acids Research, 46(D1), D1150–D1156. https://doi.org/10.1093/nar/gkx954
Yang, Z., Zhang, T., Han, S., Kusumanchi, P., Huda, N., Jiang, Y., & Liangpunsakul, S. (2021). Long noncoding RNA H19 – a new player in the pathogenesis of liver diseases. In Translational Research (Vol. 230, pp. 139–150). Mosby Inc. https://doi.org/10.1016/j.trsl.2020.11.010
Zhang, T., Hu, Y., Wu, X., Ma, R., Jiang, Q., & Wang, Y. (2016). Identifying Liver Cancer-Related Enhancer SNPs by Integrating GWAS and Histone Modification ChIP-seq Data. BioMed Research International, 2016. https://doi.org/10.1155/2016/2395341
Zhu, Y. X., Li, C. H., Li, G., Feng, H., Xia, T., Wong, C. H., Fung, F. K. C., Tong, J. H. M., To, K. F., Chen, R., & Chen, Y. (2020). LLGL1 Regulates Gemcitabine Resistance by Modulating the ERK-SP1-OSMR Pathway in Pancreatic Ductal Adenocarcinoma. CMGH, 10(4), 811–828. https://doi.org/10.1016/j.jcmgh.2020.06.009